 |
 |
 |
| |
Treatment outcome and resistance analysis in HCV genotype 1 patients previously exposed to TMC435 monotherapy and re-treated with TMC435 in combination with PegIFNa-2a/ribavirin
|
| |
| |
Reported by Jules Levin EASL 2011 March 30-Apr 2 Berlin Germany
O Lenz,1 J de Bruijne,2 L Vijgen,1 T Verbinnen,1 C Weegink,2 H van Marck,1 I Vandenbroucke,1 M Peeters,1 G De Smedt,1 K Simmen,1 G Fanning,1 G Picchio,3 H Reesink2
1Tibotec, Beerse, Belgium; 2Academic Medical Center, Amsterdam, The Netherlands; 3Tibotec Inc., Yardley, PA, USA

Introduction
· TMC435 is a once daily (QD) oral NS3/4A protease inhibitor currently
in Phase III clinical development for the treatment of hepatitis C virus
(HCV) infection.
· TMC435 is an investigational potent and selective inhibitor of the NS3/4A
protease, with an in vitro 50% effective concentration (EC50) of 8 nM
(6 ng/mL) in a genotype 1b replicon cell line.1
· Findings from Phase I and II studies have shown that TMC435 is well
tolerated, with promising efficacy reported in combination with pegylated
interferon a-2a/ribavirin (PegIFNa-2a/RBV).2-6
· In vitro resistance studies identified changes at amino acid positions
43, 80, 155, 156 and 168 within the NS3 protease domain that confer
variable degrees of reduced susceptibility to TMC435.7
· Median/mean trough TMC435 concentrations observed following
doses of 75 mg and 200 mg QD were approximately 25-50 and
350-950 fold higher, respectively, than the EC50 values obtained
in the genotype 1b-replicon cell line (6 ng/mL).2,8,9
· In a Phase I study (TMC435-C101; NCT00938899), six HCV
genotype 1-infected patients who failed previous IFN-based therapy
received TMC435 200 mg QD for 5 days.2 Approximately
1.5 years later 5/6 patients (1 patient died 1 year after study
TMC435-C101; unrelated to HCV therapy) were enrolled into Cohort 5
of the Phase IIa Optimal Protease inhibitor Enhancement of Response
to TherApy (OPERA)-1 study (TMC435-C201; NCT00561353) and
were re-treated with TMC435 (200 mg QD) in combination with
PegIFNa-2a/RBV for 28 days followed by PegIFNa-2a/RBV up to Week 48.3
· Here we report the final clinical outcomes for the 5 patients included
in the TMC435-C101 study who were re-treated in TMC435-C201, and
present a detailed genotypic and phenotypic analysis of HCV genotype 1
isolates obtained from these patients.
METHODS
· Plasma samples were collected during the TMC435-C101 and TMC435-C201 studies and HCV ribonucleic acid (RNA) levels were assessed with a
Roche COBAS® TaqMan® HCV/HPS v2.0 assay. This assay has a lower limit of
quantification (LLOQ) of <25 IU/mL. HCV RNA results below the LLOQ are
reported as either <25 IU/mL detectable or <25 IU/mL undetectable.
· HCV RNA was extracted from plasma samples and the NS3/4A protease
region was sequenced using standard population sequencing, if HCV RNA
was >100 IU/mL. In addition, selected samples were analysed using clonal
single genome sequencing (20-80 clones per sample) or were subjected
to massive parallel pyrosequencing (ultra-deep sequencing) using the 454
life science platform (GS-FLX, Roche Applied Science).
· For phenotypic analysis, selected mutations or patient-derived NS3
protease regions were introduced into a genotype 1b replicon backbone.
The antiviral activity of TMC435 was tested and compared with the
parental replicon in a standard transient replicon assay.
RESULTS
Antiviral Activity and Efficacy
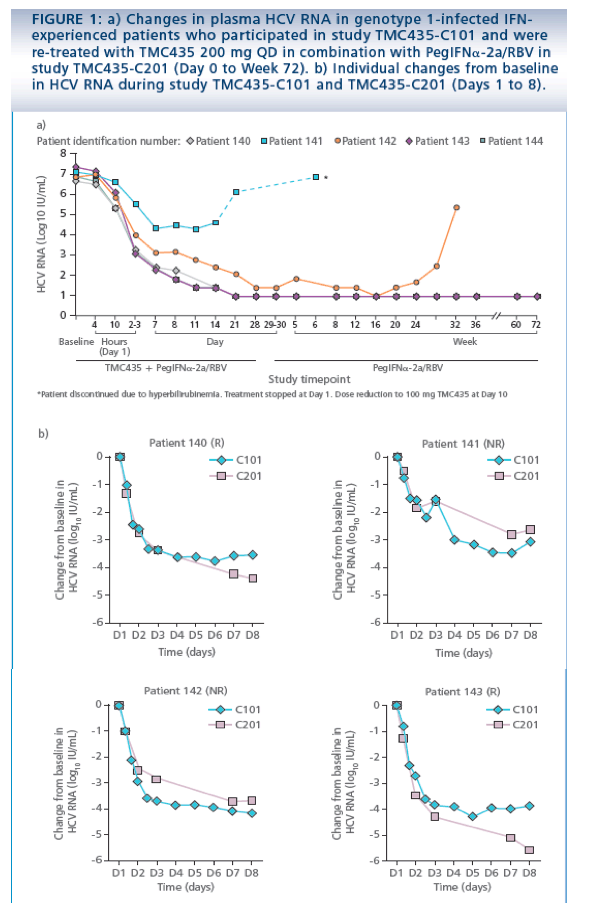
· At Day 6 in study TMC435-C101, a consistent and rapid decline in
plasma HCV RNA was observed in all six patients, with a maximal median
reduction from baseline of 3.9 log10 IU/mL.2
· Of the five patients who were re-treated in study TMC435-C201,3 one
(patient 141, previous non-responder) stopped all treatment at Day 14 due
to an adverse event (AE) (increase in bilirubin level). Three of the remaining
four patients (one non-responder and two patients with viral relapse) who
completed the 28-day TMC435 in combination with PegIFNa-2a/RBV
treatment had undetectable HCV RNA levels at Day 28 and ultimately
achieved a sustained virologic response (SVR). One patient (patient 142)
achieved HCV RNA <25 IU/mL detectable at Day 28, followed by a viral
breakthrough during PegIFNa-2a/RBV therapy at Week 28 (Figure 1a).
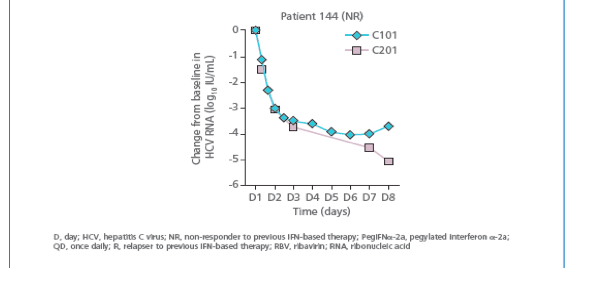
· Overall changes in HCV RNA during the first week of therapy were similar
between study TMC435-C101 and TMC435-C201. In two patients, the
HCV RNA reduction from baseline was slower in TMC435-C201 versus
study TMC435-C101: in patient 141 at Day 3 (-2.9 versus -3.7 log10 IU/mL)
and patient 142 at Day 7 (-2.8 versus -3.5 log10 IU/mL) (Figure 1b). This
difference could be attributable to persistent virus and/or suboptimal
response to PegIFNa-2a/RBV in study C201.
Genotypic/Phenotypic Changes in the HCV NS3 Protease
Region During Study TMC435-C101 and TMC435-C201
· Mutations at one or more of the amino acid positions 80, 155, 156 and
168, which were absent at baseline, were detected in all subjects as early as
on Day 3 of study TMC435-C101 using population sequencing (Figure 2a).
· Population sequencing and phenotypic analysis of the HCV isolates
obtained at baseline of TMC435-C201 suggested that the dominant
viral variants returned to the characteristics observed prior to study
TMC435-C101 in all 5 patients.
· Mutations S122R plus R155K (patient 142), and Q80R plus D168E plus
F169L (patient 141), were detected in TMC435-C201 during treatment.
· The in vitro activity of TMC435 against the treatment emerging NS3
variants identified revealed fold changes in EC50 values (FC) >100
in all patients in study TMC435-C101 and in patient 141 and 142 in
TMC435-C201 (Figure 2b).
FIGURE 2: a) Mutations detected in the NS3 protease domain defined as changes from reference sequence (Con1 for genotype 1b and H77 for genotype 1a). b) Fold changes in EC50 values compared with wild-type assessed in a transient replicon assay using chimeric replicons harbouring the NS3 protease region derived from patient samples.
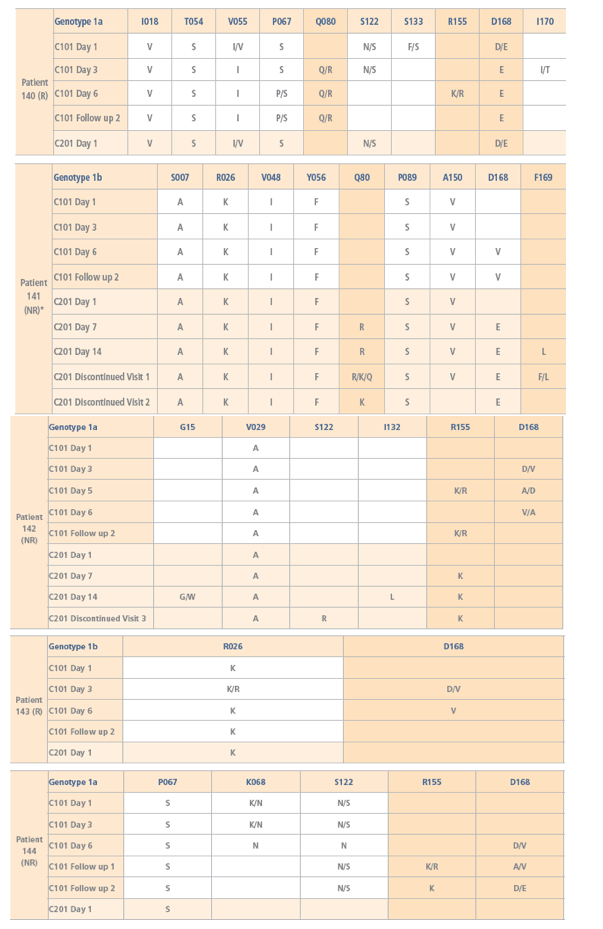
*Patient discontinued due to hyperbilirubinemia. Mutations T40A, S91T/A and L153I were found in all HCV genotype 1a patients at all time points analysed; the mutation R26K was found in all HCV genotype 1b patients at all time points analysed; A156A/V was present in patient 143 at Day 4.
NR, non-responder to previous IFN-based therapy; R, relapser to previous IFN-based therapy. Day 1 = Baseline; C101 follow-up 1 = 2 weeks after end of dosing; C101 follow-up 2 = 4 weeks after end of dosing. Missing values were below the lower limit of the assay
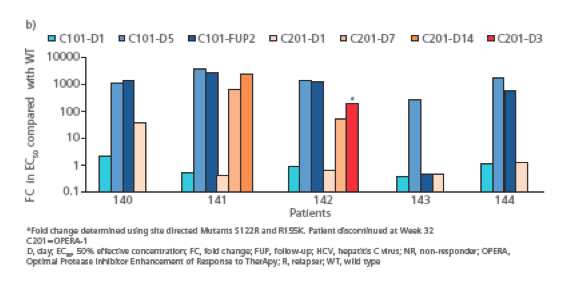
*Fold change determined using site directed Mutants S122R and R155K. Patient discontinued at Week 32 C201=OPERA-1
D, day; EC50, 50% effective concentration; FC, fold change; FUP, follow-up; HCV, hepatitis C virus; NR, non-responder; OPERA, Optimal Protease inhibitor Enhancement of Response to TherApy; R, relapser; WT, wild type
Presence of Viral Variants at Baseline in Studies TMC435-C101 and TMC435-C201 based on Deep Sequencing
· No additional NS3 mutations at key binding pocket residues (80, 155, 156 and 168) were detected at baseline of the TMC435-C101 study using ultra-deep sequencing (D168D/E was also detected by population sequencing).
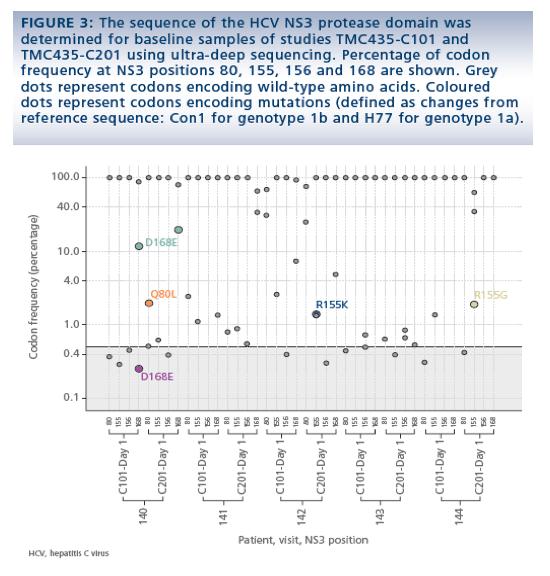
· At baseline of the TMC435-C201 study, mutations at key binding positions were observed at low frequency (<2%) in 3 patients: Q80L, R155G and R155K, in patients 140, 144 and 142, respectively (Figure 3).
- Patient 142 showed a slower initial viral decline in TMC435-C201 compared with TMC435-C101, while patients 140 and 144 showed a comparable viral decline in both studies.
Single Genome (Clonal) Sequencing of NS3 Protease Domain
· In addition to single mutations, variants encoding 2 or 3 amino acid
changes on the same genome were observed (Figure 4).
· In general, when introduced into the replicon, most of the single
mutations detected resulted in a lower fold change in EC50 value
compared with replicons with 2 or 3 mutations (data not shown).
Evolution of Amino Acid Changes over Time
Based on Deep Sequencing
· In patient 141, ultra-deep sequencing detected a Q80R and a D168E
mutation four weeks after the end of dosing for study TMC435-C101,
which were no longer detectable at baseline of the TMC435-C201 study.
These two mutations emerged as major variants upon re-exposure to
TMC435 in the TMC435-C201 study (Figure 5a). This patient also showed
a slower initial viral decline in TMC435-C201 compared with in
TMC435-C101.
· In patient 142, ultra-deep sequencing detected variant R155K persisting at
low frequency (<2%) at baseline of the TMC435-C201 study. This mutation
became detectable by population sequencing at Day 7 and 14 of the study
while HCV RNA was still declining. An S122R and an R155K mutation
became detectable by population and 454 sequencing at the time of viral
breakthrough (Week 28) (Figure 5b).
FIGURE 4: NS3 protease domain was amplified, cloned into standard
cloning vectors and sequenced to determine linkage of mutations.
Mutations at position 80, 155, 156, 168 and 169 were considered for
this analysis. Frequency of clones is represented.
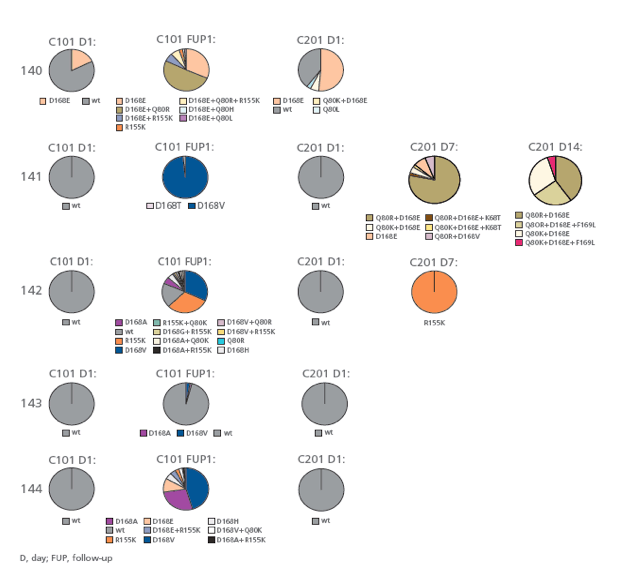
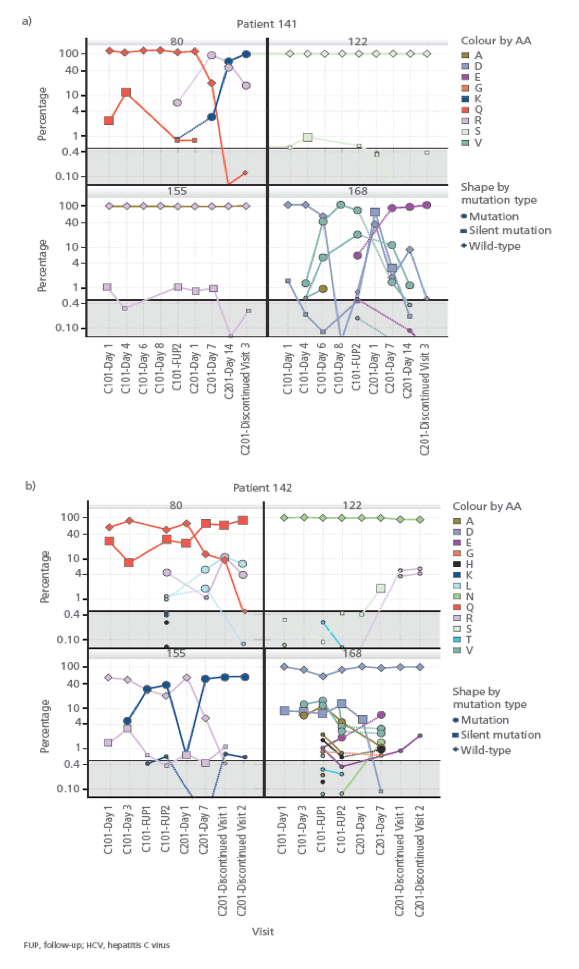
FIGURE 5: The sequence of the HCV NS3 protease domain was analysed from multiple samples using ultra-deep sequencing. Changes of codon frequency over time for NS3 at positions 80, 122, 155 and 168 are shown for two patients a) 141 b) 142.
REFERENCES
1. Lin TI et al. Antimicrob Agents Chemother 2009; 53: 1377-1385.
2. Reesink HW et al. Gastroenterology 2010; 138: 913-921.
3. Reesink H et al. Poster presented at the 60th American Association for the Study of Liver Diseases (AASLD) meeting, Boston, MA, USA, October 30 - November 3, 2009.
4. Marcellin P et al. Poster presented at the 44th Annual Meeting of European Association for the Study of the Liver (EASL), Copenhagen, Denmark, 22-26 April, 2009.
5. Manns M et al. Presented at the 44th Annual Meeting of European Association for the Study of the Liver (EASL), Copenhagen, Denmark, 22-26 April, 2009.
6. Fried MW, et al. Presented at the 61st AASLD Meeting, Boston, MA, USA, October 29 - November 2, 2010.
7. Lenz O et al. Presented at the International Symposium on Hepatitis C Virus and Related Viruses, San Antonio, TX, USA, 5-9 October 2008.
8. Sekar V et al. Presented at the 45th Annual Meeting of the European Association for the Study of the Liver (EASL), Vienna, Austria, 14-18 April, 2010.
9. Data on file. Tibotec Inc. 2010.
|
| |
|
 |
 |
|
|