 |
 |
 |
| |
HBV DNA & ALT Predict HCC in HBV+
|
| |
| |
Reported by Jules Levin
DDW, Washington DC, May 19-24, 2007
"...These results indicate that ongoing viral replication leads to biochemical abnormalities and eventual liver damage. This would suggest that viral load at a single point in time (in CHB subjects aged 30-65) captures the eventual risk of ALT elevation as well as that of future HCC risk..."
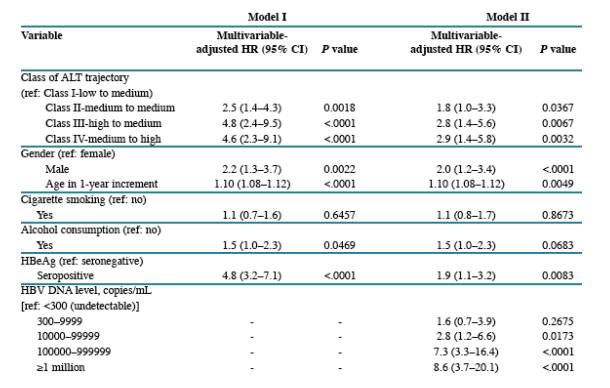
In Table 1: HBV DNA >1 million was associated with the highest risk for medium & high ALT levels (3 to 4 times higher compared to HBV DNA 100,000 to 1 million).
In Table 4: HBV DNA >1 million copies/ml increased HCC risk, associated with Hazard Ratio of 8.6 (3.7-20.1) (p <.0001) compared to HBV DNA <300 c/ml (undetectable). HBV DNA 100,000 to 1 million associated with Hazard Ratio of 7.3 (3.3-16.4) (p <.0001). HBV DNA 10,000 to 100,000 associated with Hazard Ratio of 2.8 (1.2-6.6) (p 0.0173). HBV DNA 300-10,000 associated with Hazard Ratio of 1.6 (0.7-3.9) (p 0.2675).
"Changes in Serum Alanine Aminotransferase (ALT) Level Using A Trajectory Model and Risk of Hepatocellular Carcinoma (HCC) in Chronic Hepatitis B (CHB): THE R.E.V. E. A. L.- HBV Study"
C-F Chen1,2, M-H Lee1, W-C Lee1, C-L Jen1,3, H-I Yang1,3, UH Iloeje4, J Su4, S-N Lu5, S-L You1,3, C-J Chen1,3
1National Taiwan University, Taipei, Taiwan; 2Department of Health
Administration, HungKuang University, Taichung, Taiwan; 3Genomics Research Center, Academia Sinica, Taipei, Taiwan;
4Bristol-Myers Squibb Pharmaceutical Research Institute, Wallingford, CT, USA; 5Kaohsiung Chang-Gung Memorial Hospital, Kaohsiung, Taiwan
Summary
Describing ALT Trajectories over time
- Four classes of serum ALT trajectories were identified by the model
- The majority of the study subjects were in Class II and had serum ALT between 15 and 45 U/L at study entry and remained there throughout follow-up
Baseline factors associated with serum ALT change over time
- Males were more likely than females to be in the trajectory Classes II-IV compared to Class I
- Age did not appear to be associated with changes in serum ALT over time
- Alcohol consumption and HBeAg status were associated with serum ALT changes
- The baseline HBV DNAwas strongly associated with serum ALT trajectory across a biological gradient
Serum ALT change and incidence/risk of progressing to HCC
- The incidence of HCC increased with serum ALT trajectory class
(reference subjects with Class I trajectory)
- In the Cox model, change in serum ALT as measured by the serum ALT trajectory class was a strong predictor of HCC risk
- Increasing HBV DNA level at baseline was the strongest risk predictor of HCC progression even after adjusting for changes in serum ALT over time
Author Conclusions
Through our modelling, four distinct serum ALT trajectory classes were identified, and change in serum ALT level was an independent predictor of HCC risk
Serum HBV DNA at baseline was strongly associated with the serum ALT changes over time
After adjusting for the change in serum ALT over time, baseline HBV DNA remained a strong independent predictor of progressing to HCC in this cohort, and the HCC risk increased across a biological gradient
These results indicate that ongoing viral replication leads to biochemical abnormalities and eventual liver damage
This would suggest that viral load at a single point in time (in CHB subjects aged 30-65) captures the eventual risk of ALT elevation as well as that of future HCC risk
Abstract
Background and Aims: Serum ALT is a marker of hepatocyte damage in CHB. Our aims were to:
1) evaluate the relationship between ALT changes over time and HCC risk;
2) examine the baseline factors associated with ALT changes over time.
Methods: A subgroup of the R.E.V. E. A. L.- HBV cohort of HBsAg-seropositive and anti-HCVseronegative participants, with ≥5 ALT measurements, was used in the analyses. The outcome was new HCC cases. We defined low, medium, and high ALT as <20 U/L, ≥20 U/L but <45 U/L, and ≥45 U/L, respectively. Asemi-parametric group-based modelling was used to determine major classes of ALT trajectories. Associations between baseline risk factors and the ALT trajectories were examined using polytomous logistic regression. Cox proportional hazards modelling was used to analyze the associations between the ALT trajectory classes and HCC risk.
Results: A total of 1712 subjects with 61 new HCC cases contributed 21,225 person-years of follow-up.
Four ALT trajectory classes were identified:
Class I, persistently low (n = 551);
Class II, low-to-medium (n = 808);
Class III, high-to-medium (n = 99);
Class IV, medium-to-high (n = 254).
Of the baseline risk factors, gender, age, HBeAg status, serum HBV DNA levels, and alcohol consumption were significantly associated with the ALT trajectory.
With Class I subjects as the reference group, the adjusted hazard ratio [HRadj (95% CI)] of developing HCC was 4.0 (1.2-13.4) for Class II subjects; 4.6 (1.1-18.9) for Class III subjects; and 7.4 (2.1-25.9) for Class IV subjects after adjustment for gender, age, HBeAg status, HBV DNA levels, cigarette smoking, and alcohol consumption.
With HBV DNA <300 copies/mL as reference, the HRadj ( 95% CI) of developing HCC associated with baseline HBV DNA level was 1.8 (0.5-7.4) for HBV DNA 300 to <104 copies/mL (10,000); 4.0 (1.1-14.8) for HBV DNA ≥ 104 to <105 copies/mL (100,000); 7.1 (2-25.6) for HBV DNA ≥ 105 to <106 (1 million); and 9.6 (2.5-36.2) for HBV DNA ≥ 106 copies/mL.
Conclusions: Serum ALT changes over time are a strong independent predictor of HCC. Baseline HBV DNA was a significant predictor of the ALT trajectory over time and remained an important risk factor for HCC development after adjusting for the ALT trajectories.
Introduction
The risk of disease progression in CHB is linked to several viral, biochemical, and immunologic markers
In our published analyses,1,2 baseline serum ALT was not statistically significant as an independent predictor of liver cirrhosis and HCC. Those analyses were limited by not accounting for changes in serum ALT over time
Serum ALT level is known to correlate with the level of hepatocyte damage in a variety of liver diseases, as well as with eventual mortality in the general population3
The primary objectives of these analyses were:
- Describe the latent trajectory classes of serum ALT changes over time in the
R.E.V.E.A.L-HBV study cohort
- Examine the baseline factors associated with serum ALT changes over time
- Evaluate the relationship between serum ALT changes over time and risk of progressing to HCC adjusted for other variables including baseline serum HBV DNA level
Study Design
As previously described, the R.E.V.E.A.L.-HBV study is a population-based prospective cohort study with a mean follow-up of 11 years
A subgroup of this study cohort contributed to these analyses
Of the 3,653 anti-HCV-negative, HBsAg-positive subjects with adequate serum sample for HBV DNA testing, 3,054 had follow-up serum ALT levels beyond the baseline measurement. These subjects were enrolled in these analyses
There were 132 incident cases of HCC in this subpopulation.
Analyses:
- HBV DNA of cohort entry serum samples by PCR (Roche Diagnostics Co.,
Indianapolis, IN)
- HBsAg and HBeAg by radioimmunoassay (Abbott Laboratories, North Chicago, IL)
- Anti-HCV antibody by 2nd generation ELISA kits (Abbott Laboratories, North
Chicago, IL)
- ALT by serum chemistry autoanalyser (Model 736, Hitachi Co., Tokyo, Japan) using commercial reagents (bioMérieux, Marcy l'Etoile, France)
Study Population/Methods
- Complete description of the case ascertainment for the HCC has been described1
- Semiparametric group-based modelling was used to determine major classes of serum ALT trajectories (Objective 1)
- The group-based trajectory model was designed to identify clusters of individuals following similar progressions of repeated measurements over time4
- Model selection was based on the Bayesian Information Criterion (BIC) as a measure of goodness-of-fit5
- According to the graphical summary of the group-based trajectory model, we defined low, medium, and high ALT as <15 U/L, ≥15 U/L but <45 U/L, and >45 U/L, respectively (Note: the ALT cut point was adjusted between the abstract and the final analyses, explaining the difference in cut points.)
- Associations between baseline risk factors and the serum ALT trajectories were examined using polytomous logistic regression (Objective 2)
- Cox proportional hazards modelling was used to analyze the association between the serum ALT trajectory classes and progression to HCC
- The statistical software SAS (SAS Institute Inc., Cary, NC) was used to obtain the corresponding estimates and figures
Figure 1: Study Subject Flow Chart
89,293 volunteers invited to participate in 1991-1992 (47,000 men, 42,000 women. 23,820 enrolled in study: 11,000 men, 11,000 women. 4,155 HBsAg positive; 3,851 with HBV DNA test of enrollment serum sample; 3.054 with adequate ALT follow-up measurements used in this study (1,979 men, 1,075 wimen) (132 newly diagnosed with HCC cases).
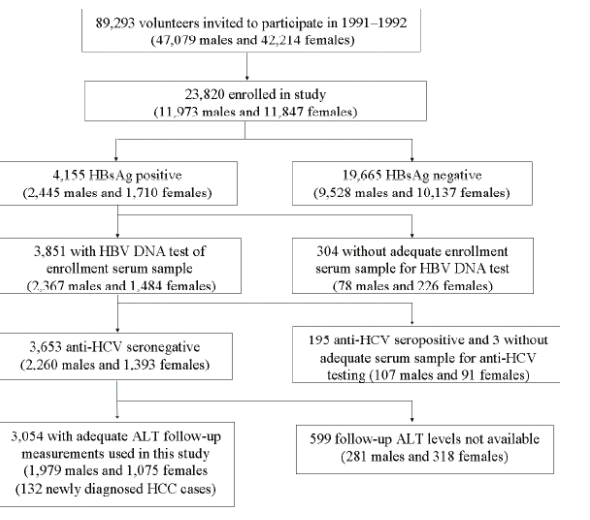
Table 1. Baseline Demographics (Whole Sub-Cohort and the Four
Serum ALT Classes Determined by the Model)
HBV DNA >1 million was associated with the highest risk for medium & high ALT levels (3 to 4 times higher compared to HBV DNA 100,000 to 1 million). HBV DNA between 300 to 1 million were associated with similar risk levels for ALT elevations, although risk for ALT elevation was greater then when HBV DNA <300 c/ml.
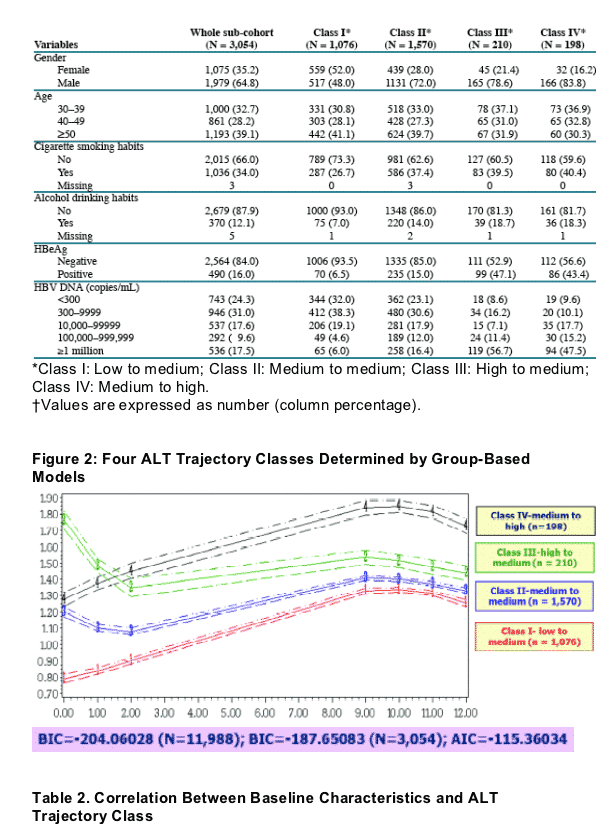
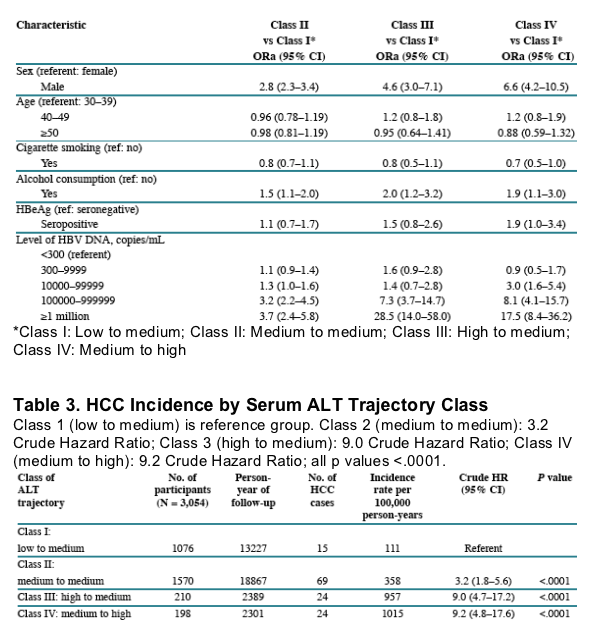
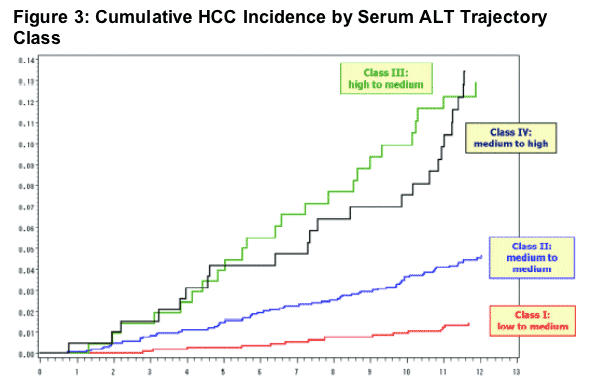
Table 4. HCC Incidence by Serum ALT Trajectory Class
HBV DNA >1 million copies/ml increased HCC risk, associated with Hazard Ratio of 8.6 (3.7-20.1) (p <.0001) compared to HBV DNA <300 c/ml (undetectable). HBV DNA 100,000 to 1 million associated with Hazard Ratio of 7.3 (3.3-16.4) (p <.0001). HBV DNA 10,000 to 100,000 associated with Hazard Ratio of 2.8 (1.2-6.6) (p 0.0173). HBV DNA 300-10,000 associated with Hazard Ratio of 1.6 (0.7-3.9) (p 0.2675).
References
1. Chen CJ, Yang HI, Su J et al. Risk of hepatocellular carcinoma across a biological gradient of serum hepatitis B virus DNA level. JAMA. 2006;295:65-73.
2. Iloeje UH, Yang HI, Su J, et al. Predicting liver cirrhosis risk based on the level of circulating hepatitis B viral load. Gastroenterology. 2006;130:678-686.
3. Kim WR, Benson JT, Therneau TM, et al. Serum aminotransferase concentrations and risk of mortality in a community population. Hepatology. 2006;44(suppl 1):655A. Abstract 1251.
4. Jones BL, Nagin DS, Roeder K. A SAS procedure based on mixture models for estimating developmental trajectories. Sociol Meth Res. 2001;29:374-393.
5. Schwarz G. Estimating the dimension of a model. Ann Stat. 1978;6:461-464.
Financial Disclosures
Chuen-Fei Chen: Nothing to declare
Mei-Hsuan Lee: Nothing to declare
Wen-Chung Lee: Nothing to declare
Chin-Lan Jen: Nothing to declare
Hwai-I Yang: Nothing to declare
Uchenna H. Iloeje: Bristol-Myers Squibb employee
Jun Su: Bristol-Myers Squibb employee
Sheng-Nan Lu: Nothing to declare
San-Lin You: Nothing to declare
Chien-Jen Chen: Research support: Bristol-Myers Squibb
|
| |
|
 |
 |
|
|