 |
 |
 |
| |
Genotypic Analysis of Patients with Evaluable HBV DNA After 1 Year of Telbivudine Therapy in the GLOBE Registration Trial
|
| |
| |
Reported by Jules Levin
DDW, May 22, 2007, Wash DC
M. Seifer1; A. Patty1; C. Chapron1; L. van Doorn2; B. Belanger1; N. A. Brown1;
D. N. Standring1
1. Idenix Pharmaceuticals, Cambridge, MA, USA.
2. DDL Diagnostic Services, Voorburg, Netherlands.
ABSTRACT
Introduction: The safety and efficacy of telbivudine was evaluated in the international GLOBE trial of 1367 patients with chronic hepatitis B. Here we report the genotypic analysis at week 48 of 165 telbivudine patients with HBV of >1,000 copies per mL.
Methods: HBV genomes were amplified from sera of study patients by RT PCR. The entire polymerase RT domain was sequenced to determine HBV genotypes at screen. For 165 telbivudine patients with HBV DNA >1,000 copies per mL at week 48 or 52, sera were re-sequenced at week 48 (or post virologic breakthrough) to identify genotypic mutations. These included all patients who had met previous virologic breakthrough criteria at week 48.
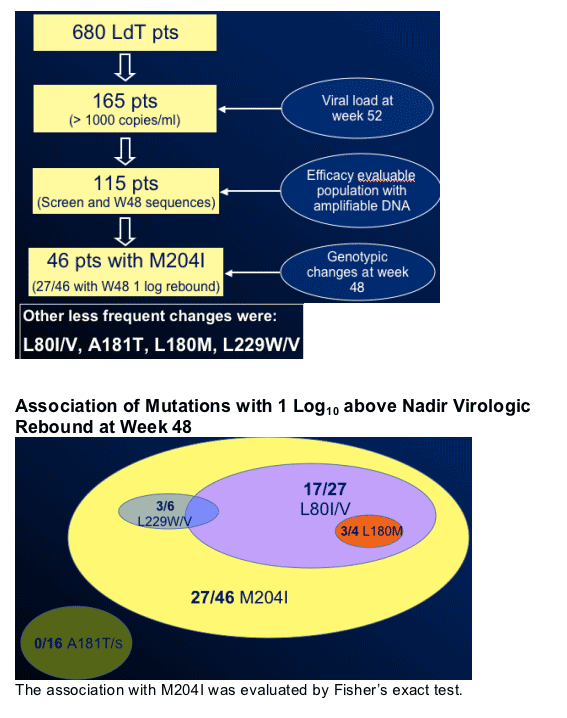
Results: Sequences were obtained from 115/165 evaluable patients (with amplifiable HBV DNA and at least 16 weeks on therapy). RtM204I was the most frequent genotypic change seen (in 46/115 patients, including 9 mixed mutants) and is the only mutant to date that has been causally linked to genotypic/phenotypic telbivudine resistance and virologic rebound: at week 48, M204I was detected in 28 of 32 telbivudine patients experiencing a 1 log above nadir breakthrough (Standring et al, DDW 2007). Other genotypic changes among the 115 patients included rtL80 (n=26), rtL180 (4), and rtL229 (6). These were only associated with virologic rebound when rtM204I was present. There was no evidence for the L180M/M204 resistance pattern for telbivudine.
The genotypic analysis revealed rtA181T/S substitutions (n=15/1) that did not co-segregate with M204I or virologic rebound. These mutants result in a prematurely terminated S protein, likely ablating viable virus production. The mutation was absent in the 5 A181T patients who went on to year 2 breakthrough (3 with M204I, 2 with wild-type M204). Thus, the biological relevance and role in resistance of this variant is obscure.
Conclusions: In the large GLOBE study, telbivudine primarily elicited M204I genotypic changes (n= 46). M204I, with or without secondary mutation (primarily L80I), is the signature telbivudine mutation associated with virologic rebound. No L180M/M204V-based resistance/changes were seen for telbivudine.
SLIDE PRESENTATION
Genotypic Changes After 1 Year of Telbivudine Therapy in the GLOBE Registration Trial
Maria Seifer, April Patty, Christopher Chapron, Leen-Jan van Doorn, Bruce Belanger, Nathaniel A Brown, David N Standring
AUTHOR SUMMARY
Telbivudine Resistance Profile -
M204I is telbivudine signature resistance mutation
Close HBV monitoring for lack of early virologic response (i.e. at 24 weeks) identifies those patients at risk for developing resistance to telbivudine monotherapy
In vitro cross resistance studies predict telbivudine resistance can be salvaged by add-on of nucleotides (ADV or TNV)*
*Supported by preliminary clinical data from Heathcote et al (Poster 1776) (NATAP report coming)
Telbivudine (LdT)
Specific inhibitor of HBV polymerase
-- Not active in vitro against HIV or other viruses or against cellular DNA polymerases
Favorable preclinical toxicology:
-- Non-mutagenic, no mitochondrial toxicity in vitro
-- Non-carcinogenic, no teratogenicity or embryofetal toxicities
-- Pregnancy Category B designation in US label
Once daily oral dosing indicated by PK
-- Consistent absorption, no food effect
First and second year of Phase III GLOBE comparative trial
-- Greater antiviral efficacy vs lamivudine in HBeAg+ and HBeAg- patients, with less resistance and comparable safety
Telbivudine Clinical Resistance Profile:
Resistance Determination Strategy
Population sequencing
-- Ability to detect novel mutants
-- Commonly used in HBV pivotal trials to determine clinical resistance profile
Determination of the entire polymerase RT domain
-- All 344 amino acids covering all known resistance mutations
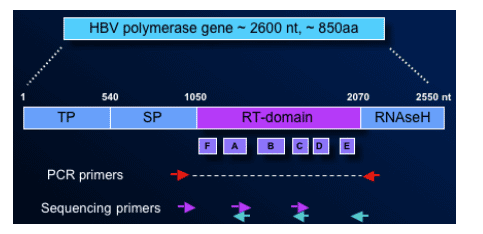
Blinded samples sequenced by independent, 3rd party vendor (Delft Diagnostic Laboratories - Voorburg, Netherlands)
Telbivudine Clinical Resistance Profile:
Resistance Determination Strategy
Breakthrough definition criteria:
-- [Per protocol: serum HBV DNA returns to >5 log10, or within 1 log10 of baseline]
-- Confirmed 1 log10 above nadir HBV DNA rebound
Analysis presented on:
-- GLOBE pts with 1 log10 above nadir viral rebound
-- Telbivudine pts with serum HBV DNA
>1,000 copies per mL

Telbivudine
A simple pattern of resistance based on M204I mutant
(no M204V)
Typically a 2- to 4-fold lower clinical resistance rate than Lam
No novel or unique telbivudine resistance mutation found

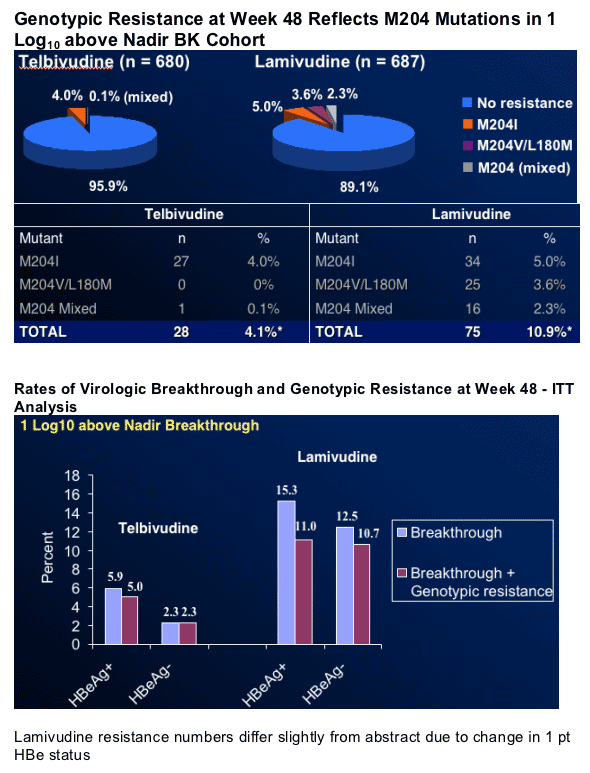
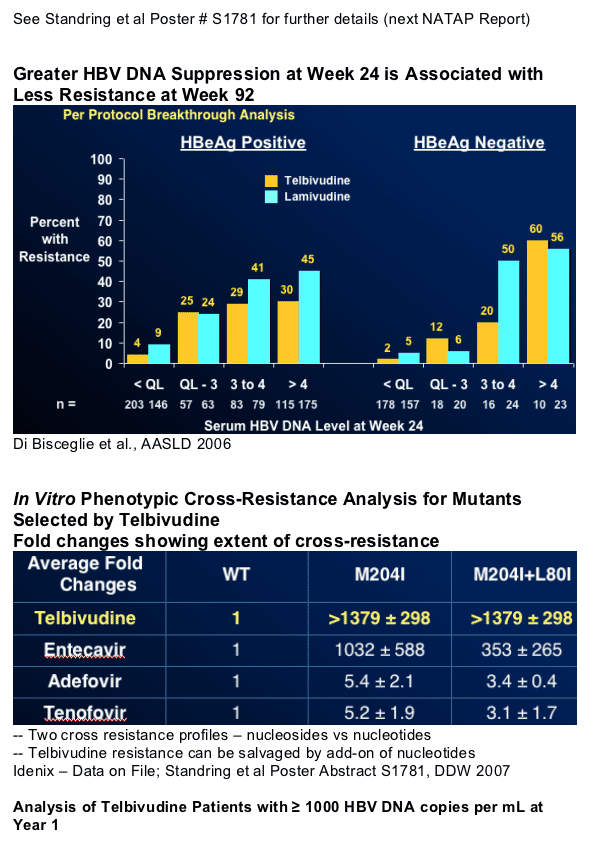
M204I: Telbivudine Signature Mutation
Mutation M204I is the primary pathway to breakthrough and genotypic resistance for telbivudine
The clinical resistance profile validates predictions from in vitro models
-- Telbivudine is active against M204V single mutation
-- M204V mutation precedes M204V/L180M lamivudine resistance
Other mutations that occur with M204I are most likely secondary/compensatory mutants
-- codon L80 mutants seen in 2/3 of M204I pts
-- occasional changes at codons L180 or L229
A181T Mutant: No Clear Link to Telbivudine Resistance
A181T is infrequent in GLOBE
-- Half of the observed mutations are mixed
A181T introduces stop codon into the S gene (codon 172)
A181T mutant is not detected in any patient with viral rebound at Year 1 or Year 2
A181T is not associated with M204I
|
| |
|
 |
 |
|
|