| |
Viral hepatitis: HCV compartmentalization in HCC: driver, passenger or both? - Diminished viral replication and compartmentalization of hepatitis C virus in hepatocellular carcinoma tissue
|
| |
| |
Download the PDF here
Download the PDF here
"The high frequency of synonymous substitutions suggested that the changes in the viral variants observed were not a result of selection. When the degree of HCV RNA replication restriction was further analysed, patients with HCC could be divided into those with either a <2 log or >2 log reduction in HCV RNA levels between the nontumorous and tumorous tissue. The degree of change in the HCV quasispecies diversity was significantly greater in those with more HCV replication restriction (>2 log versus <2 log HCV RNA reduction, P = 0.0257). Interestingly, when malignant cell proliferation was assessed by staining for the enzyme MIB1 in a subset of the HCC samples, those with a >2 log reduction in HCV RNA level demonstrated a greater degree of malignant hepatocyte proliferation than those with a <2 log reduction. These data suggest that there is a higher rate of tumour cell proliferation in patients who have increased HCV quasispecies diversity and restriction of HCV replication, which might have implications for HCC pathogenesis."
EASL: Development of Hepatocellular Carcinoma in HCV Cirrhotic Patients Treated with Direct Acting Antivirals - (04/25/16)
----------------------------
Nature Reviews Gastroenterology & Hepatology | News and Views
Viral hepatitis: HCV compartmentalization in HCC: driver, passenger or both?
Jacinta A. Holmes and Raymond T. Chung
Gastroenterology Division, Massachusetts General Hospital, Harvard Medical School, 55 Fruit Street, Boston, Massachusetts 02114, USA.
Published online
23 March 2016
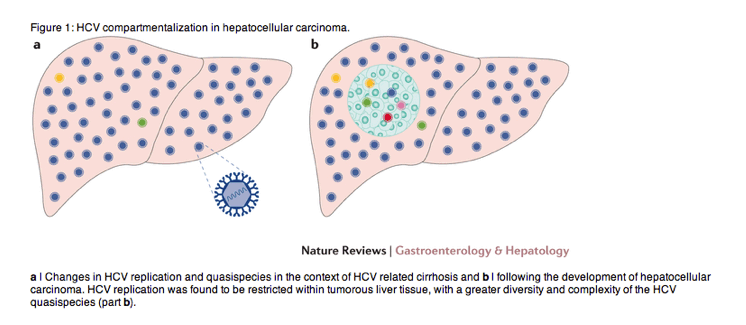
Hepatocellular carcinoma is associated with HCV infection but the underlying interplay between virus and tumour remains to be elucidated. Now, Harouaka et al. report that in patients with HCV-related cirrhosis, HCV replication is restricted within liver tissue originating from hepatocellular carcinoma, with an associated increase in the diversity and complexity of the HCV quasi species.
Refers to Harouaka, D. et al. Diminished viral replication and compartmentalization of hepatitis C virus in hepatocellular carcinoma tissue. Proc. Natl Acad. Sci. USA 113, 1375-1380 (2016) [see below article]
The incidence of hepatocellular carcinoma (HCC) has been sharply rising globally with the number of incident cases tripling over the past 15 years, largely driven by HCV infection1. Chronic inflammation from HCV infection results in progressive liver damage and fibrosis, leading to the complications of cirrhosis, HCC and end-stage liver failure2. HCC still carries a poor prognosis, and is one of only a few cancers with a rising mortality. Early diagnosis of HCC is critical to improve long-term survival, as early-stage disease can be amenable to curative therapies. However, it is currently difficult to predict which patients infected with HCV will develop HCC, and therefore all those with advanced fibrosis are enrolled in screening programmes for HCC. In their new paper3, Harouaka and colleagues report that HCV replication is severely restricted within HCC tissue compared with the surrounding nontumorous liver, and that the genetic diversity (genetic distance between viral variants) and complexity (number of viral variants) of HCV also differs substantially between tumorous and nontumorous tissue. Taken together, these novel findings suggest there is compartmentalization of the virus in cirrhotic livers from HCV-infected patients with HCC (Fig. 1).
The precise role of HCV in HCC pathogenesis has not been completely defined, although both direct and indirect roles have been identified4. HCV exists as quasispecies of viral variants in liver and serum that are strongly influenced by host immune pressure5. This pressure allows the quasispecies to evolve over time, developing escape mechanisms that permit viral persistence with the hypothesis that, as the disease progresses, HCV diversity decreases due to less immune pressure. Thus, there has been much interest in evaluating HCV diversity, complexity and replication in both serum and liver tissue from patients infected with HCV to gain further understanding of HCV-related HCC pathogenesis. However, efforts have been limited by the lack of adequate experimental models and access to high-quality samples from multiple areas of the liver from the same patient.
In the latest study, Harouaka et al.3 showed that the contrast in HCV replication between tumorous and nontumorous tissue was not related to changes in microRNA-122 expression (a well-described host factor required for HCV replication6) and, moreover, circulating HCV RNA levels were found to be comparable between patients with (n = 8) and without (n = 4) HCC. Differences in HCV quasispecies diversity were not observed in areas of nontumorous tissue adjacent to the HCC, the distant peripheral non-HCC tissue, or between the right and left lobes of the non-HCC cirrhotic control livers. Arbitrary variability of HCV RNA levels was, therefore, excluded as an explanation for the observed findings. The authors then assessed whether changes in quasispecies could be attributed to natural selection by assessing the proportion of synonymous (non-coding) and nonsynonymous (coding) viral RNA substitutions. The high frequency of synonymous substitutions suggested that the changes in the viral variants observed were not a result of selection. When the degree of HCV RNA replication restriction was further analysed, patients with HCC could be divided into those with either a <2 log or >2 log reduction in HCV RNA levels between the nontumorous and tumorous tissue. The degree of change in the HCV quasispecies diversity was significantly greater in those with more HCV replication restriction (>2 log versus <2 log HCV RNA reduction, P = 0.0257). Interestingly, when malignant cell proliferation was assessed by staining for the enzyme MIB1 in a subset of the HCC samples, those with a >2 log reduction in HCV RNA level demonstrated a greater degree of malignant hepatocyte proliferation than those with a <2 log reduction. These data suggest that there is a higher rate of tumour cell proliferation in patients who have increased HCV quasispecies diversity and restriction of HCV replication, which might have implications for HCC pathogenesis.
One of the advantages of the latest study is that multidirectional sampling of HCC tissue (up to five samples from the tumour) and surrounding non-HCC liver tissue (up to 17 samples in total, averaging 13 samples per patient) was performed in HCV-infected patients with cirrhosis and HCC, thereby overcoming the issue of inadequate liver sampling. Paired sera were available allowing comparison of the characteristics of circulating HCV to intrahepatic HCV, and HCV-infected patients with cirrhosis but without HCC were included as controls, with samples taken from the right and left lobes of the liver (four samples per patient). This comprehensive sampling strategy permitted a more global assessment of HCV RNA levels and quasispecies within both HCC tissue and surrounding nontumorous tissue than many previous studies. However, it is still difficult to conclude from data taken at a single time point whether the changes in HCV replication and the HCV quasispecies are driving tumorigenesis, or if these changes are induced by the tumour milieu. Although serial tissue sampling over time or the availability of precancerous lesions would help shed light on this question, both are practically challenging to obtain. Nonetheless, some intriguing data support the concept that perhaps the observed findings are both the cause and effect of HCC.
The interferon-induced double-stranded RNA-activated protein kinase (EIF2AK2, also known as PKR) recognizes double-stranded RNA and phosphorylates eukaryotic translation initiation factor 2 subunit 1 (also known as eukaryotic initiation factor-2 or eIF-2α), which has been shown to block HCV replication by inhibiting initiation of HCV translation7. Levels of PKR and eIF-2α are also significantly elevated in HCC liver tissue compared with surrounding non-tumorous tissue (P = 0.001)8, suggesting that the host response to the tumour might induce factors that alter localized immune pressure on HCV. Interestingly, overexpression of PKR has been associated with reduced HCV RNA levels in HCC tissue9, suggesting that overexpressed PKR remains functionally active in HCC. PKR also upregulated the JNK1 and ERK1/3 pathways, which led to increased HCC cell proliferation in the context of HCV infection. HCV-driven PKR expression might therefore act both to control localized HCV levels as well as to promote tumour growth in response to HCV infection. Such a model is amenable to testing and suggests that PKR could be an interesting therapeutic target.
As the incidence of HCC continues to rapidly increase, largely driven by the ageing HCV population10, its poor prognosis and extremely limited therapies contribute to the observation that HCC is now the fastest rising cause of global cancer-related mortality1. We currently remain unable to predict with precision which patients will develop HCC. Thus, because patients with advanced fibrosis or cirrhosis are at risk, it is still recommended that all of these patients be enrolled in screening programmes for HCC. Although tempting to speculate that local HCV RNA concentrations or quasispecies variations within the liver could foreshadow HCC, the utility of such a tool would be limited given the need for extensive tissue sampling and the complexity of quasispecies analysis. Also, it remains to be seen whether these viral compartment changes are only found in the context of established HCC. As such, identification of alternative biomarkers for future HCC risk would be extremely useful, even in the face of a successful HCV cure now achievable with contemporary antiviral therapy. In this regard, use of gene expression signatures from HCV-infected cirrhotic tissue could be important for prognosis. The study by Harouaka et al.3 sheds light on the events that attend HCV-related tumorigenesis, but further studies are needed to identify possible mediators of these findings.
--------------------------------
Proc. Natl Acad. Sci. USA (2016)
Diminished viral replication and compartmentalization of hepatitis C virus in hepatocellular carcinoma tissue
Djamila Harouakaa, Ronald E. Englea, Kurt Wollenbergb, Giacomo Diazc, Ashley B. Ticea, Fausto Zambonid,
Sugantha Govindarajane, Harvey Alterf,1, David E. Kleinerg, and Patrizia Farcia,1
aHepatic Pathogenesis Section, Laboratory of Infectious Diseases, National Institute of Allergy and Infectious Diseases, National Institutes of Health,
Bethesda, MD 20892; bBioinformatics and Computational Biosciences Branch, Office of Cyber Infrastructure and Computational Biology, National Institute
of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, MD 20892
Significance
Hepatocellular carcinoma (HCC) associated with hepatitis C virus (HCV) infection is the fastest-rising cause of cancer-related death in the United States. The level of intratumor HCV replication and the molecular interactions between virus and tumor remain elusive, however. Here we demonstrate that the ability of HCV to replicate in HCC is severely hampered despite unchanged miR122 expression. Surprisingly, we found that livers containing HCC harbor a more diverse viral population than that seen in cirrhotic livers without HCC. Tracking of individual variants demonstrated changes in quasispecies distribution between tumor and nontumorous areas, suggesting viral compartmentalization within the tumor. These insights into the interplay between HCV and HCC call for further investigation of whether malignant hepatocytes express or lack factors that restrict HCV entry or negatively affect viral replication.
Abstract
Analysis of hepatitis C virus (HCV) replication and quasispecies distribution within the tumor of patients with HCV-associated hepatocellular carcinoma (HCC) can provide insight into the role of HCV in hepatocarcinogenesis and, conversely, the effect of HCC on the HCV lifecycle. In a comprehensive study of serum and multiple liver specimens from patients with HCC who underwent liver transplantation, we found a sharp and significant decrease in HCV RNA in the tumor compared with surrounding nontumorous tissues, but found no differences in multiple areas of control non-HCC cirrhotic livers. Diminished HCV replication was not associated with changes in miR-122 expression. HCV genetic diversity was significantly higher in livers containing HCC compared with control non-HCC cirrhotic livers. Tracking of individual variants demonstrated changes in the viral population between tumorous and nontumorous areas, the extent of which correlated with the decline in HCV RNA, suggesting HCV compartmentalization within the tumor. In contrast, compartmentalization was not observed between nontumorous areas and serum, or in controls between different areas of the cirrhotic liver or between liver and serum. Our findings indicate that HCV replication within the tumor is restricted and compartmentalized, suggesting segregation of specific viral variants in malignant hepatocytes.
Hepatitis C virus (HCV) is a hepatotropic, single-stranded RNA virus that replicates in the cytoplasm of hepatocytes and does not integrate into the host genome (1). HCV circulates in vivo as a dynamic distribution of closely related viral variants that are commonly referred to as "quasispecies" (2). Such diversity confers a remarkable advantage to the virus under host selective constraints (3 ⇓-5). One of the most important features of HCV is its extraordinary ability to persist in up to 80% of infected individuals (6). Approximately 20-30% of chronic HCV carriers develop cirrhosis and its long-term sequelae, including hepatic decompensation and hepatocellular carcinoma (HCC), leading to orthotopic liver transplantation (OLT) or liver-related death (6). The incidence of HCC in the United States has more than tripled over the past 3 decades (7), an alarming trend due primarily, if not exclusively, to HCV infection (8).
Although epidemiologic evidence has linked chronic HCV infection to a significantly elevated risk of developing HCC (9), the mechanisms whereby HCV promotes hepatocarcinogenesis remain to be elucidated (10). Whether HCV elicits liver cancer indirectly, through chronic inflammation, fibrosis, and continuous liver regeneration (11), or directly, through the expression of tumor-promoting viral proteins, in a manner analogous to other oncogenic viruses, such as human papillomaviruses and Epstein-Barr virus (12, 13), remains unknown. The major challenges in defining the role of HCV in the pathogenesis of HCC include the inherent limitations of available experimental systems, the low number of infected hepatocytes, the low level of HCV antigen expression, and the limited availability and size of infected liver samples (14).
Insights into the levels of HCV replication within the tumor of patients with HCV-associated HCC may shed light on the role of HCV in hepatocarcinogenesis. This remains controversial, however, in part because of the difficulty in obtaining multiple paired liver specimens from the tumor and adjacent nontumorous tissue. Although some previous investigations have shown no differences in the presence and levels of HCV RNA between the tumor and nontumorous liver tissues (15 ⇓-17), others have reported low to undetectable levels of HCV RNA within the tumor (18 ⇓-20). Several reports have indicated that miR-122 is an essential cellular host factor for HCV replication (21, 22). Initial studies performed in liver cancer, regardless of the etiology, have shown a reduced expression of miR-122 (23, 24), whereas recent reports have demonstrated that its expression in HCV-associated HCC is maintained (25, 26) or even increased (27). Thus, the central questions of whether HCV actively replicates in malignant hepatocytes, its relationship with miR-122 expression, and whether viral replication is directly involved in hepatocarcinogenesis remain to be fully elucidated. A related issue is whether changes in viral replication within the tumor are associated with selection of specific viral variants; however, very limited information is available on the composition of the HCV quasispecies and the possible compartmentalization between the tumor and the surrounding nontumorous areas or serum from the same individuals with HCV-associated HCC (20, 28 ⇓-30).
To investigate the role of HCV in hepatocarcinogenesis, we took advantage of a unique collection of multiple liver specimens from patients with HCV-associated HCC who underwent OLT or partial hepatectomy to simultaneously study the level of viral replication, its correlation with the intrahepatic expression of miR-122, and the composition and distribution of the viral population both within and outside the tumor.
Results
Patients.
The demographic, clinical, virologic, and histopathological features of the 12 patients with HCV-associated HCC (n = 8) or non-HCC cirrhosis (n = 4) included in this study are presented in Table S1. The mean age of the two groups was similar; all patients except one were male (92%) and all but one were infected with HCV genotype 1. The grade of tumor differentiation was G2 in four patients and G3 in four patients; in all cases, HCC was surrounded by a cirrhotic liver.
HCV RNA Levels in Patients with HCC and Controls with Non-HCC Cirrhosis.
An average of 13 liver specimens from each patient with HCC and 4 liver specimens from each patient with non-HCC cirrhosis were tested for the presence and levels of intrahepatic HCV RNA by real-time PCR (Fig. 1 A and B). A sharp and significant drop in viral RNA levels was observed in all of the patients with HCC when perilesional tissue was compared with tissue inside the tumor margin (Fig. 1C), with the lowest concentration in the center of the tumor (area A). In contrast, HCV RNA levels were significantly higher and comparable among the three nontumorous areas, C, D, and E (Fig. 1C), as well as in four different regions of control livers, spanning the right and left lobes obtained from the four patients with non-HCC cirrhosis (Fig. 1D). In individual patients, the extent of the drop in mean HCV RNA levels between the surrounding nontumorous areas (C, D, and E) and the tumor (areas A and B) ranged between 0.89 and 3.15 logs. Five patients (patients 1-5) showed a <2-log drop in HCV RNA (HCC pattern 1), whereas three patients (patients 6-8) had a >2-log drop (HCC pattern 2) (Fig. 1E). In contrast, no changes in HCV RNA levels were detected between the right and left lobes of livers from the controls with non-HCC cirrhosis (Fig. 1F). The HCC and non-HCC patients had comparable levels of HCV in serum (Table S1 and Fig. S1 A and B). Thus, our study demonstrates that HCC is associated with a severe restriction of intrahepatic HCV replication.
Expression of miR-122 in Patients with HCC and Controls with Non-HCC Cirrhosis.
To investigate the mechanisms underlying the significant reduction in HCV replication within HCC tissues, we examined the expression of miR-122 in multiple areas of the livers containing HCC, because this miRNA has been shown to be essential for HCV replication (31). Interestingly, we found no differences in miR-122 expression among the different areas of the liver containing HCC (Fig. 2A), indicating that the significant reduction in HCV RNA within the tumor is not related to changes in miR-122 expression. Similar levels of miR-122 were also seen in different areas of the liver in controls with non-HCC cirrhosis (Fig. 2B).
Genetic Diversity, Complexity, and Distribution of HCV Quasispecies in Liver Compartments and Serum of Patients with HCC and Non-HCC Cirrhosis.
We next studied the genetic distance between the different variants (i.e., genetic diversity) and the number of viral strains (i.e., genetic complexity) of the HCV quasispecies, both within and outside the hypervariable region 1 (HVR1) of the E1/E2 region in HCC patients divided according to the extent of HCV RNA decrease (patterns 1 and 2) and in controls with non-HCC cirrhosis (Fig. S2A). We found a statistically significant decrease in HCV RNA in HCC by comparing the tumor with the surrounding nontumorous tissue (P < 0.01 for pattern 1; P < 0.000001 for pattern 2). We also found a significant difference between HCV RNA levels in HCC pattern 1 and pattern 2 within the tumor (P = 0.04), but not in the surrounding nontumorous tissues. HCV RNA levels were also significantly higher in non-HCC cirrhosis than in the tumor (P < 0.0001 for pattern 1; P < 0.000001 for pattern 2), whereas there were no differences in serum HCV RNA levels between the two groups of patients.
The genetic diversity, as measured by Hamming distance, was significantly higher in the livers of patients with HCC compared with those of controls with non-HCC cirrhosis both within and outside the HVR1, irrespective of the liver compartment analyzed (P < 0.0007 in all comparisons) (Figs. S2B and S3). Genetic diversity was also higher in the serum of patients with HCC, both within and outside the HVR1 (P = 0.016 and P = 0.004, respectively) (Figs. S2B and S3). Among the HCC patients, the degree of genetic diversity within HVR1 was significantly higher in the center of the tumor in patients with pattern 2 compared with those with pattern 1 (P = 0.0257) (Fig. S2B).
The number of viral variants, as assessed by HVR1 sequences, was significantly higher in the liver of patients with HCC pattern 2 than in controls with non-HCC cirrhosis (P = 0.002) or in patients with HCC pattern 1 (P = 0.029), whereas no differences were detected between patients with HCC pattern 1 and these controls (Fig. S2C). A significantly higher number of variants in serum samples was also observed in patients with HCC pattern 2 compared with controls with non-HCC cirrhosis (P = 0.025).
Tracking of individual viral variants in multiple liver compartments and serum demonstrated two different patterns of viral quasispecies distribution, which correlated with the drop in HCV RNA levels from the surrounding nontumorous areas to the tumor. In patients with HCC pattern 1 (patients 1-5; Fig. 1E), who had the smallest drop in HCV RNA (<2.0 log), the predominant variant within the tumor was either dominant or codominant in all liver compartments, as well as in serum (Fig. 3 B and D). Interestingly, the center of the tumor contained unique minor variants that were not detected in the periphery of the tumor, but lacked some of the variants that started to appear in the periphery of the tumor and further expanded to the surrounding nontumorous areas (Fig. 3 B and D). Patients with HCC pattern 2 (patients 6-8; Fig. 1E), who had the most dramatic drop in HCV RNA (>2.0 logs), exhibited a more complex HCV quasispecies with greater changes in the distribution of the viral variants (Fig. 3 F and H). A distinctive feature of this group was the behavior of the dominant strain found in the center of the tumor, which coexisted with other major variants and lost dominance both at the periphery of the tumor and in the surrounding nontumorous areas (Fig. 3 F and H). The virus population in the nontumorous liver compartments was similar in the patients with HCC pattern 1 and HCC pattern 2 and mirrored the quasispecies distribution in serum (Fig. 3 B, D, F, and H).
In contrast to the changes in quasispecies distribution observed in the eight patients with HCC, analysis of multiple liver specimens and serum of the four control patients who never developed HCC showed a different pattern that was consistent across all of the liver areas analyzed, characterized by the presence of a dominant strain, representing 65-95% of the viral population, along with some minor variants (Fig. 3 J and L). The dominant strain and one or two minor variants were shared by all of the liver compartments, as well as by the viral population circulating in serum (Fig. 3 J and L).
Malignant Cell Proliferation in Patients with HCC.
We next investigated the rate of proliferation of malignant hepatocytes in the patients with HCC pattern 1 and pattern 2. Formalin-fixed tissue obtained from the center of the tumor of four patients with HCC was available for staining for the cell proliferation marker MIB1. Two of these four patients (patients 3 and 4) exhibited pattern 1, with a <2-log drop in HCV RNA, and the other two (patients 7 and 8) exhibited pattern 2, with a >2-log drop in HCV RNA (Fig. 1E). The percentage of tumor nuclei positive for MIB1 was assessed using an ocular grid at 20×. For each patient, more than 1,000 tumor nuclei were assessed (mean, 1,640; range, 1,130-2,350). Remarkably, we found that patients with the lowest level of HCV RNA showed the highest proportion of proliferating malignant cells. The percentage of MIB1 positivity was 0.55% and 4.0% in pattern 1 patients 3 and 4, respectively (Fig. 4 A and B), compared with 12.4% and 15.8% in pattern 2 patients 7 and 8 (Fig. 4 C and D). These data suggest that the high viral diversity documented within the tumor of patients with HCC pattern 2, who had the most significant decrease in HCV RNA in the liver (>2 logs), is associated with a higher rate of cell proliferation compared with patients with HCC pattern 1.
HCV Compartmentalization and Selection Analysis in Patients with HCC and in Controls with Non-HCC Cirrhosis.
To determine whether the differences in HCV quasispecies distribution between the tumorous and nontumorous tissues of patients with HCC reflects compartmentalization of viral variants in different areas of the liver, we used the Mantel test (32).
Compartmentalization was defined by a significant difference in virus populations (quantified as pairwise genetic distance among the sequences) between different areas of the liver or between the liver and serum. Remarkably, the viral population differed significantly between the tumor and nontumorous tissues in all eight patients with HCC (Table 1). In more than 50% of the patients (four of seven), the viral population detected in the tumor also differed from that circulating in serum (Table 1). In contrast, no significant differences were found between the viral population detected in nontumorous tissues (areas C, D, and E) and the variants circulating in serum (Table 1), indicating that the viral population circulating in serum is most likely produced in the surrounding nontumorous areas, which represents the largest area of the liver.
In contrast, in the controls with non-HCC cirrhosis, our analysis revealed no evidence of HCV compartmentalization between the right and left lobes, or between liver compartments and serum (Table 1). Thus, our data indicate that HCC is associated with viral compartmentalization.
We next investigated whether the different patterns of HCV quasispecies distribution and HCV compartmentalization between the tumorous and nontumorous tissues of patients with HCC could be attributed to positive selection. We estimated the mean number of nonsynonymous and synonymous substitutions per nonsynonymous and synonymous site, respectively, across the E1/HVR1/E2 region within the tumor and the nontumorous tissues, as well as among different areas of the control non-HCC cirrhotic livers, and in the serum of both groups of patients. An excess of nonsynonymous substitutions with respect to synonymous substitutions was considered to be indicative of positive selection. Overall, the mean number of synonymous substitutions per site was greater than that of nonsynonymous substitutions per site across the E1/HVR1/E2 region, both within and outside the tumor, as well as in serum (Fig. S4A). A similar pattern was seen in cirrhotic patients (Fig. S4B). These results suggest that HCV compartmentalization and the distribution of viral variants within the tumor is not the result of a selective host immune pressure.
Discussion
To the best of our knowledge, this is the first study in which the levels of HCV replication and the distribution of the viral quasispecies were comprehensively investigated in serum and multiple compartments of individual livers containing HCC and compared with levels in control livers with non-HCC cirrhosis. Although the number of patients that could be included in this intensive study was limited, our patients were well characterized and devoid of confounding factors such as viral coinfections, i.v. drug use, alcohol abuse, or antiviral treatment. All were Caucasian, and all but one was infected with HCV genotype 1. Analysis of HCV RNA in up to 17 specimens from each liver containing HCC provided conclusive evidence that the tumor is associated with restricted levels of viral replication.
Previous studies performed by in situ PCR (33, 34), fluorescence microscopy (35), or laser-capture microdissection followed by RT-PCR (36, 37) indicated that the distribution of HCV RNA in the liver is focal and that the percentage of infected hepatocytes is low (never >35% of the total population). However, the focal distribution of HCV-infected cells (37) cannot explain the restricted viral replication that we detected exclusively within the tumor. Our analysis, extended to the entire liver containing HCC, clearly demonstrates that the marked decline in HCV RNA occurs exclusively within the tumor. Thus, the drop in viral RNA is highly specific to the malignant hepatocytes and cannot be related to sampling variability or to the presence of a mixture of malignant and normal hepatocytes, given that all liver specimens with a mixed-cell population were excluded from the analysis.
Consistent with these findings, and in agreement with previous studies (38 ⇓-40), reduced HCV RNA levels were not detected in any of the surrounding nontumorous areas or in different regions of the right and left lobes of livers from patients with cirrhosis who never developed HCC, indicating that nontumorous liver tissue can efficiently sustain HCV replication regardless of the presence of a tumor. Thus, the decline in HCV RNA documented exclusively within the tumor would have been missed by analyzing only the levels of viremia or routine biopsy specimens of nontumor tissue.
The fact that HCV does not replicate well in malignant hepatocytes is consistent with the inability or limited efficiency of HCV to grow in hepatoma cell lines in vitro (1), suggesting that malignant hepatocytes express factors, or more likely have lost the expression of factors, that may restrict viral entry or negatively affect viral replication. Interestingly, our analysis of multiple paired liver specimens obtained from the tumor and surrounding nontumorous tissue showed no differences in miR-122 expression, indicating that the reduced levels of HCV RNA in the tumor were not the result of a reduction in miR-122 expression. The level of miR-122 in HCC compared with the surrounding nontumorous tissue has been a matter of debate (23 ⇓ ⇓ ⇓-27). Our observation is consistent with the results of recent studies that showed no changes in miR-122 expression between tumorous and nontumorous tissue in HCV-associated HCC (25, 26).
Our access to a unique collection of liver and serum samples provided us with the opportunity to study the relationship between HCV RNA replication and viral quasispecies distribution within and outside the tumor. Surprisingly, we found that livers containing HCC harbor a more complex viral population with a significantly higher genetic diversity compared with cirrhotic livers without HCC. Although there was no difference in overall viral diversity between the tumor and surrounding nontumorous tissues, despite the significant drop in HCV RNA within the tumor, tracking of individual viral variants showed distinct changes in the composition and distribution of the viral quasispecies. Of note, the extent of the variation in quasispecies distribution appeared to correlate with the differential decline in HCV RNA between nontumorous and tumorous compartments; the greater the drop in HCV RNA within the tumor, the greater the shift in viral quasispecies distribution between the tumor and surrounding nontumorous tissues.
Our data also show that the differences in quasispecies distribution are not the result of differences in host selective pressure in malignant hepatocytes, as was suggested previously (20). The molecular mechanisms leading to high viral diversity within the tumor despite the significant drop in viral replication remain to be elucidated. We found a higher rate of proliferation of malignant hepatocytes in the center of the tumor in the patients with HCC pattern 2, who maintained a high degree of diversity despite the greatest drop in HCV RNA, suggesting that proliferation of neoplastic cells may play a role in maintaining a high degree of viral diversity within the tumor. This is an interesting observation that needs to be confirmed in a large series of patients.
Changes in malignant hepatocytes may lead to the expression, or lack of expression, of factors that contribute to maintaining a segregation of viral variants between tumor and nontumor compartments. Consistent with this hypothesis, Mantel test analysis demonstrated significant differences in HCV quasispecies distribution between the tumorous and nontumorous compartments, providing evidence of HCV compartmentalization between these two liver areas. In contrast, analysis of multiple liver areas in the control patients with non-HCC cirrhosis demonstrated no evidence of HCV compartmentalization within the liver or between the liver and serum, further indicating that the presence of HCV compartmentalization is unique to tumor-containing livers (20, 41). Interestingly, viral compartmentalization between plasma and extrahepatic sites, including leukocytes (42), PBMCs (43), and the brain (44), also has been reported in HCV-infected patients.
In summary, our study shows that patients with HCC harbor a more complex HCV population in the liver compared with patients with non-HCC cirrhosis. Moreover, our data provide conclusive evidence that HCC is associated with restricted HCV replication within the tumor despite unchanged miR122 expression levels, as well as with HCV compartmentalization. Whether and to what extent HCV-infected tumor cells harbor replication-competent virus remain to be defined, although this question is difficult to address owing to the lack of small animal models and efficient in vitro systems for growing primary HCV isolates. Our results provide insights into the role of HCV in hepatocarcinogenesis and may help to elucidate whether malignant hepatocytes express or lack factors that restrict viral entry or replication.
|
|
| |
| |
|
|
|