 |
 |
 |
| |
EVALUATION OF bNAb SENSITIVITY BY GENOTYPING AND PHENOTYPING FOR HIV CLINICAL TRIALS
|
| |
| |
CROI 2021 March 6-10 Reported by Jules Levin
Brian Moldt1, PC Parvangada1, Ross Martin1, Craig Pace1, Mini Balakrishnan1, Nathan Thomsen1, Herbert Kuster2, Dominique Braun2, Huldrych F. Günthard2, Sean E. Collins1, Romas Geleziunas1, Christian Callebaut1
1Gilead Sciences, Inc, Foster City, CA, USA, 2University Hospital Zurich, Zurich, Switzerland
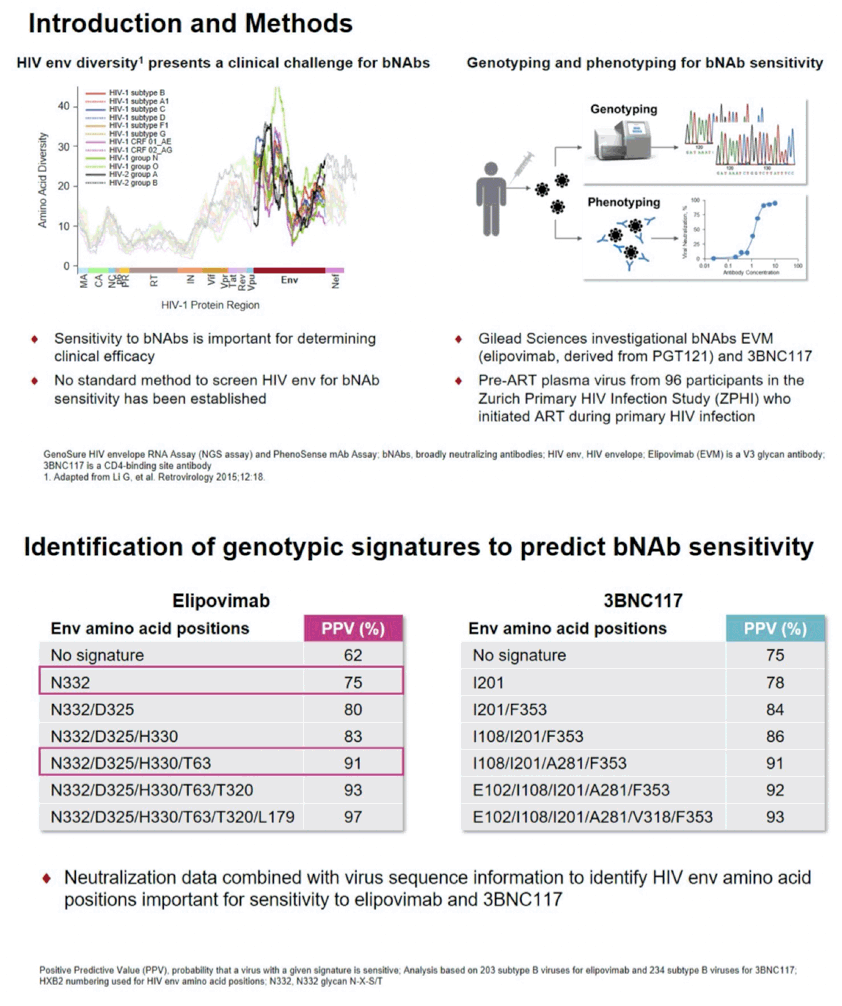
Background: HIV envelope (Env) diversity is a significant challenge for the use of broadly neutralizing antibodies (bNAbs) in HIV treatment and cure studies. Screening Env for bNAb susceptibility to select sensitive participants will be important to improve clinical efficacy, however, no standard approach has been established. Individuals who initiate ART during primary HIV infection generally have low sequence diversity and are an attractive population for early proof of concept bNAb cure-related trials. We therefore analyzed Env sequences from individuals who started ART during primary HIV infection.
Methods: Pre-ART plasma virus from 96 participants in the Zurich Primary HIV Infection Study, who initiated ART during primary HIV infection, was genotyped and phenotyped for susceptibility to the bNAbs elipovimab (EVM, formerly GS-9722) and 3BNC117. The genotypic GenoSure HIV Envelope RNA Assay and the phenotypic PhenoSense HIV nAb Assay assessments were carried out following CLIA guidelines as would be required for entry into clinical trials. For predicting bNAb susceptibility by genotyping, Env amino acid signatures for EVM and 3BNC117 sensitivity were identified from a genotypic-phenotypic correlation algorithm using a subtype B database (n=203 for EVM, N=234 for 3BNC117). A positive predictive value (PPV), was calculated for each genotypic Env signature based on phenotypic sensitivity to the bNAbs.
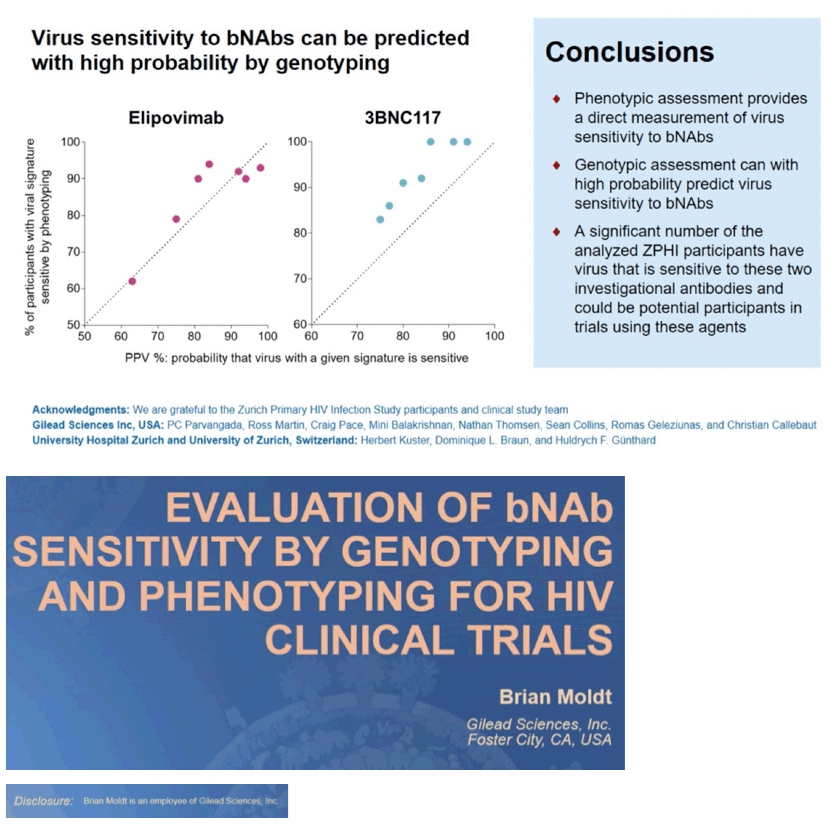
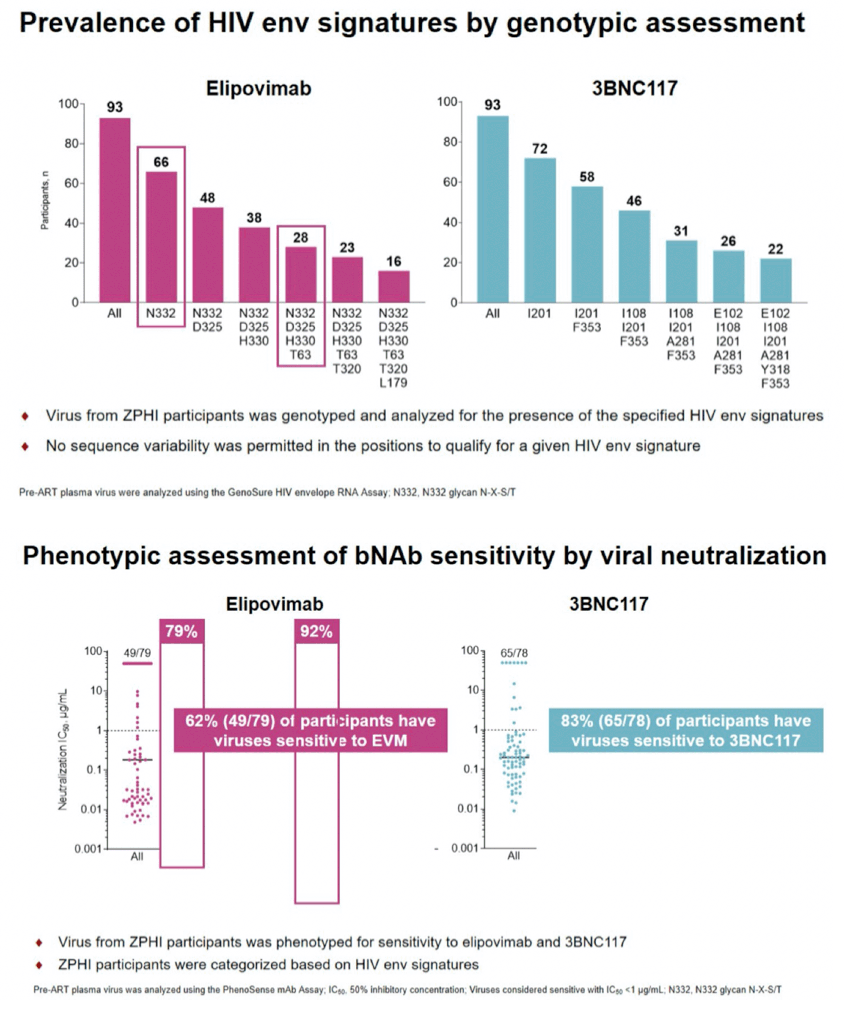
Results: Bioinformatic methods identified Env signatures with PPVs from 75% to 97% for EVM, achieving higher PPVs required more complex Env signatures. Genotyping the plasma virus and applying Env sensitivity signatures with PPVs of 75%, 83%, 91%, 97% identified 57, 32, 25 and 14 participants, respectively, out of 96 as sensitive to EVM. Plasma viruses were also evaluated for sensitivity to EVM by phenotyping. For the 57, 32, 25 and 14 participants predicted to be sensitive by genotyping, 45 (79%), 30 (94%), 23 (92%) and 13 (93%) participants, respectively, were confirmed sensitive to EVM by phenotyping. Similar analyses were performed for 3BNC117.
Conclusion: The genotypic assessment using the developed Env signatures for sensitivity appears as predictive as the direct measurement of sensitivity by phenotyping and may therefore be preferred due do turnaround time and assay simplicity. A significant number of the analyzed participants had Env sequences that are susceptible to EVM and 3BNC117 and could thus be potential candidates for trials involving these bNAbs.
|
| |
|
 |
 |
|
|