 |
 |
 |
| |
ELUCIDATING THE ORIGINS OF HIV-1 IN THE CEREBROSPINAL FLUID
|
| |
| |
CROI 2021 March 6-10 Reported by Jules Levin
Olivia D . Council1, Laura Kincer1, Sarah B. Joseph1, Susan Morgello2, Benjamin B. Gelman3, Ronald Swanstrom1
1University of North Carolina at Chapel Hill, Chapel Hill, NC, USA, 2Icahn School of Medicine at Mt Sinai, New York, NY, USA, 3University of Texas at Galveston, Galveston, TX, USA
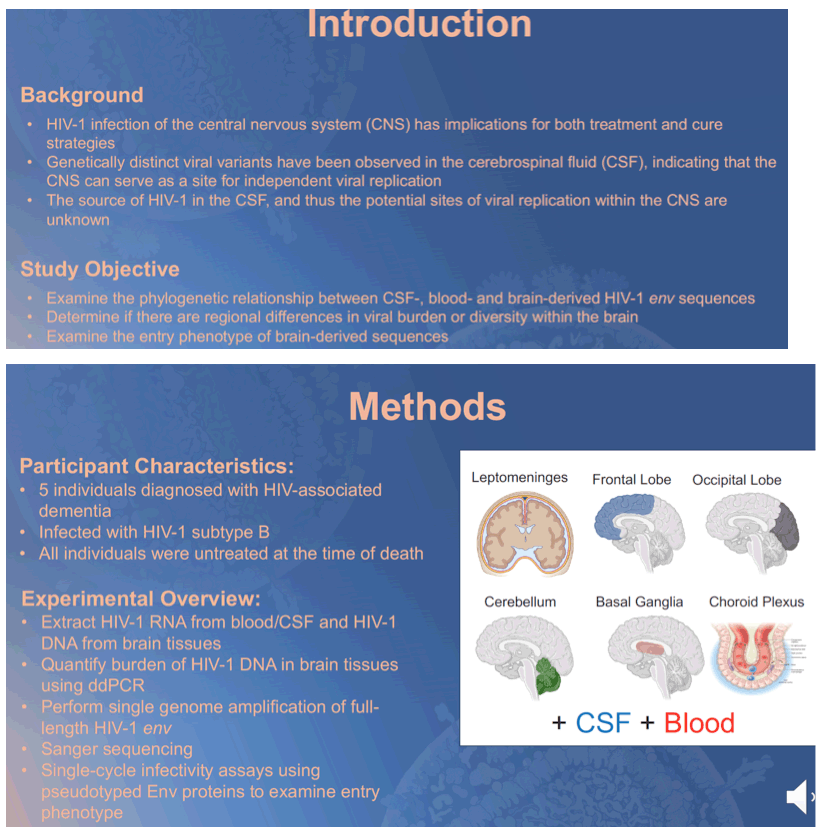
Background: HIV-1 infection of the CNS has implications for both treatment and cure strategies. Putative CNS reservoirs would likely be linked to sites of viral replication. HIV-1 RNA in the cerebrospinal fluid (CSF) can occasionally be genetically distinct from virus in the blood, indicating that the CNS can serve as a site for independent viral replication. However, the source of HIV-1 in the CSF from within the CNS is unknown and thus the sites of viral replication, and sites of potential reservoirs, remain to be determined.
Methods: Paired blood, CSF, and autopsy tissue samples from multiple brain regions were obtained from 5 participants enrolled in the NNTC. These individuals were diagnosed with HIV-associated dementia, and no participant was taking ART at the time of death. HIV-1 genomes were extracted from blood/ CSF (RNA) and brain tissues (DNA). The burden of HIV-1 DNA in various regions of the brain was quantified using ddPCR. Single genome amplification was performed on blood, CSF, and brain tissue samples in order to obtain full-length HIV-1 env sequences. Pseudotyped reporter viruses were generated from a total of 32 env genes cloned from 5 participants (22 from brain or CSF and 10 from blood plasma) and used in single-cycle entry assays to assess entry phenotype.
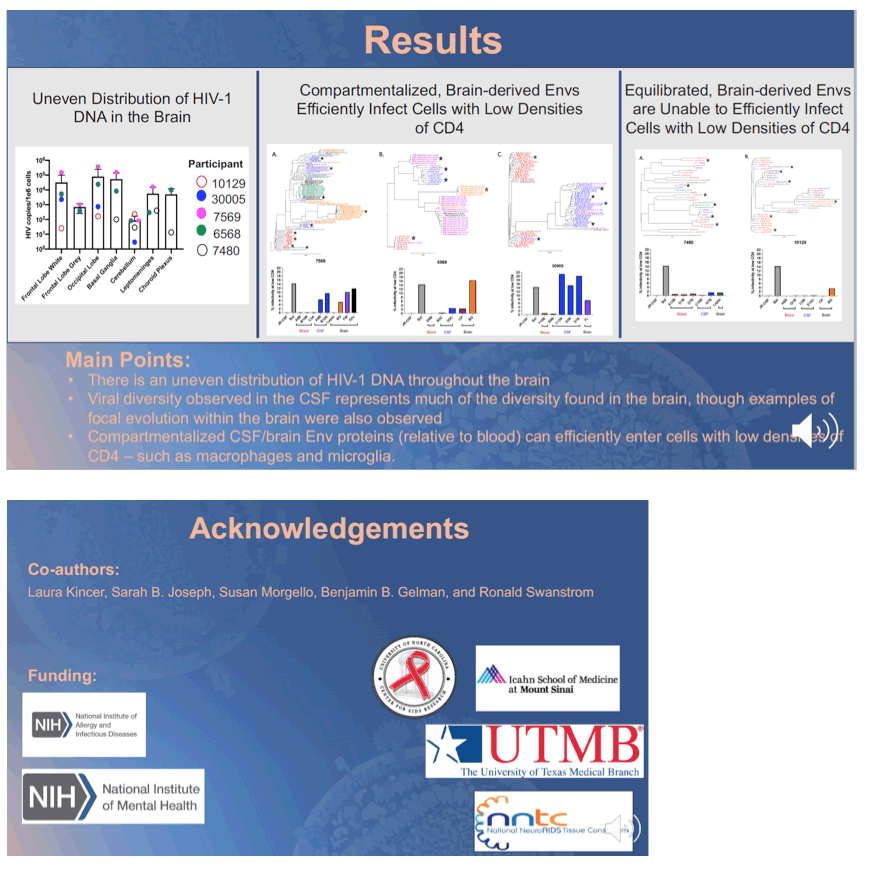
Results: In this cohort of 5 untreated people with HAD, the burden of HIV- infected cells varied across different regions of the brain, ranging from below the limit of detection to over 100,000 copies per million cells. In 3/5 participants, brain- and CSF-derived sequences were compartmentalized from blood- derived sequences. Overall, viral diversity present in the CSF represents much of the viral diversity found in the brain, but brain-specific lineages were also observed. In 2/5 participants, blood, CSF, and brain-derived sequences were intermingled, suggestive of a lack of independent viral evolution within the CNS. Compartmentalized CSF/brain Env proteins mediated efficient entry into cells expressing low densities of CD4 (M-tropic), whereas equilibrated Env proteins did not (R5 T-tropic).
Conclusion: In this small, pilot study we observed that the viral diversity present within the CSF represents much, but not all, of the viral diversity present within various regions of the brain. Indeed, viral diversity and entry phenotype can vary across different regions of the brain in the same participant. Altogether, the CSF captures the majority of the genotypic and phenotypic properties of HIV-1 in the CNS of individuals with HAD.

|
| |
|
 |
 |
|
|