 |
 |
 |
| |
NEUROSYMPTOMATIC HIV-1 CSF ESCAPE ASSOCIATES WITH REPLICATION IN CNS CD4+ T CELLS
|
| |
| |
CROI 2022 Feb 11-16
Laura Kincer1, Ameet N. Dravid2, Paola Cinque3, Mattia Trunfio4, Andrea Calcagno4, Serena S. Spudich5, Magnus Gisslen6, Richard Price7, Sarah B. Joseph1 1University of North Carolina at Chapel Hill, Chapel Hill, NC, USA, 2Poona Hospital and Research Center, Pune, India, 3San Raffaele Scientific Institute, Milan, Italy, 4University of Turin, Turin, Italy, 5Yale University, New Haven, CT, USA, 6University of Gothenburg, Gothenburg, Sweden, 7University of California San Francisco, San Francisco, CA, USA
program abstract
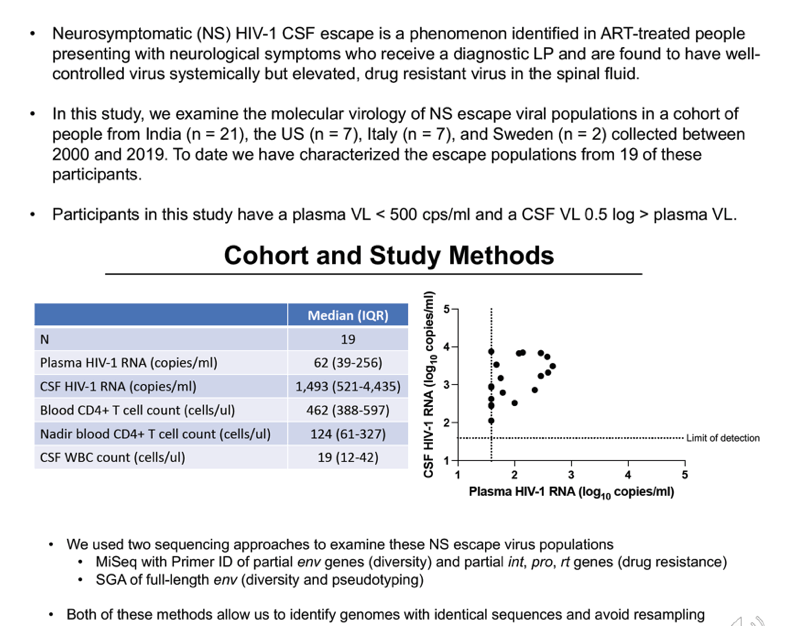
Background: Neurosymptomatic (NS) cerebrospinal fluid (CSF) escape occurs in people living with HIV-1 who present with neurologic symptoms and have elevated HIV-1 RNA in their CSF despite being on antiretroviral therapy (ART) and having undetectable or low levels of HIV-1 RNA in the blood. The cellular source of NS escape virus is unknown.
Methods: We examined viral populations in blood plasma and CSF from people with NS CSF escape (neurologic symptoms with CSF viral load (VL) >40 copies/ml, and CSF VL > plasma VL, n=36). We used single genome amplification (SGA) and/or Illumina MiSeq deep sequencing with Primer ID to asses genetic diversity in partial env sequences (V1-V3) and drug resistance in PR, RT, and IN. Full-length env genes were cloned from the CSF of 10 participants and their ability to enter cells expressing a low density of CD4 (a predictor of macrophage tropism) was assessed. Pairwise distances of partial envs were calculated.
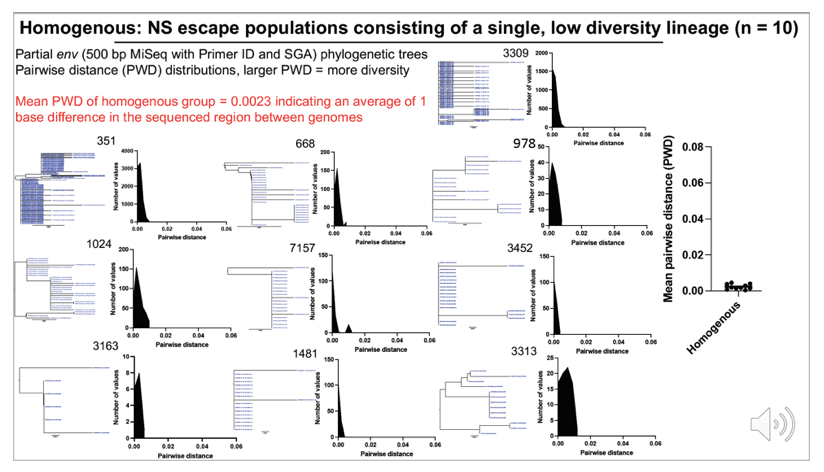
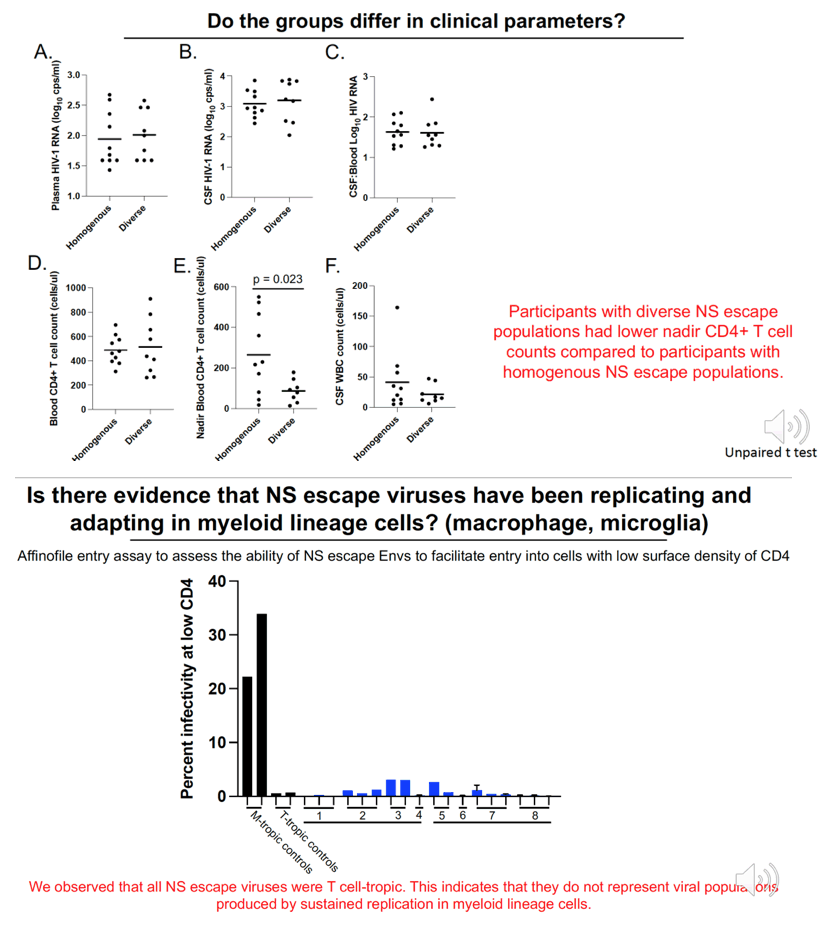
Results:Median CSF and plasma VLs were 2,400 and 140 copies/ml, respectively. The median blood CD4 count was 471 cells/ul, nadir CD4 count 98 cells/ul, and CSF white blood cells (WBC) 21 cells/ul. CSF NS escape populations either had one major lineage (47%), two major lineages (33%) or a highly diverse, recombinant population (16%). Participants with the most diverse populations also had lower nadir CD4 counts compared to those with single lineages (t test, p = 0.04). 97% of escape populations were at least partially resistant to their ART regimen and, of those, 94% saw symptoms improve after ART optimization. All escape viruses examined (N=25 from 10 participants) were T cell-tropic.
Conclusion: Observed genetic diversity, drug resistance and resolution of symptoms after ART optimization indicate that NS escape is produced by replication of partially drug resistant virus in the CNS of ART-treated participants. In addition, profound immunosuppression prior to ART was associated with escape populations with higher genetic diversity likely due to persistent replication in the CNS of these participants. The escape viruses remaining T cell-tropic despite replication and evolution in the CNS suggests that they primarily replicate in CD4+ T cells. Together these results illustrate that NS escape populations are produced by replication of partially drug resistant viruses in the CNS and suggest that evolution of escape virus is facilitated by elevated CSF WBCs and an impaired ability of the immune system to control viral replication.


|
| |
|
 |
 |
|
|